7 min read
Functional Assays: Narrowing the Field from Binders to Therapeutic Candidates
Candidate ID: How to go from hits to lead candidates
We tend to think of antibodies as being defined by their specificity and affinity to a particular antigen. Of equal importance, however, are the functional activities of an antibody upon antigen binding. All are valuable to the discovery of therapeutic antibodies, where the goal is to have a drug that can react with a specific disease target and modify its role in the disease. We describe in this post the types of assays and strategies used to identify the most promising therapeutic antibody candidates.
Binding Assays: Separating the Wheat From the Chaff
Immunoassays encompass a set of commonly used assays that are tailored towards the detection of antibody binding. They come in many formats and variations, including:
-
Enzyme-Linked Immunosorbent Assay (ELISA)
-
Flow Cytometry
-
Label-free biomolecular interaction such as biolayer interferometry (BLI) and surface plasmon resonance (SPR)
-
Newer homogeneous and multiplexed binding assays such as fluorescence energy transfer (FRET), luminescence (e.g., AlphaLISA) and electrochemiluminescence (e.g., Meso-scale Discovery)
In their simplest form, these assays can be used to identify antibody binding, eliminating weak target binders, non-binders, and antibodies with inappropriate specificity. The assays can typically be used with unpurified antibody samples (e.g., serum and supernatant) and are amenable to moderate to high-throughput screening. Most formats are inexpensive on a per sample basis; they are also quantitative, and they offer a large dynamic range.
Functional Assays: Narrowing the Field
Functional assays give you insight into an antibody’s effect on its target and its potential to be disease modifying. An antibody can function by directly modifying the activity of the target (e.g., acting as an agonist or antagonist) or by eliciting immune reactions (e.g., complement and cell-mediated cytotoxicity) through its Fc domain. Since Fc effector function can be engineered later in drug development, the direct modification of the target bio-activity is typically of greatest interest in antibody discovery, which we discuss below.
Target activity modification can be measured via a bioassay or ligand binding assay. The latter is commonly used to determine if an antibody blocks the binding of the target to its receptor. This can be done using ELISA formats for soluble antigens or using Flow Cytometry for cell-associated antigens. Cell-based assays are best for measuring antibody binding to natively expressed membrane protein targets. They can also be formatted to determine if an antibody inhibits ligand binding to the membrane protein target. An example for an antibody clone supernatant is shown below. In this assay, a fluorescently labeled ligand of the membrane protein target with strong binding to cells bearing the target is blocked by an anti-target antibody.
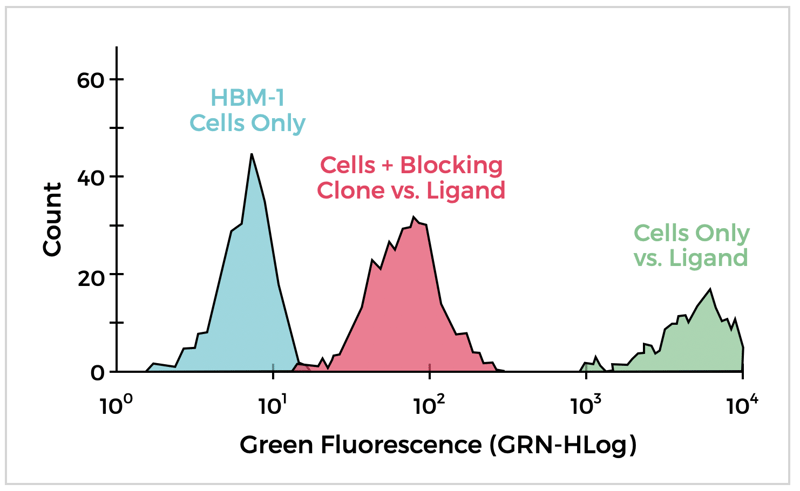
Using Flow Cytometry to Show Antibody Blocking of Ligand Binding to a Cell Membrane Receptor Protein.
Some immunoassay formats, like the one described above, may require labeling of reagents with a detector (e.g., biotin, fluorochrome, enzyme). The labels can potentially interfere with antibody or ligand binding. By contrast, label-free analysis platforms (e.g., BLI and SPR) can remedy this issue since detectors are not needed. As an example shown below and in the associated poster, a functional assay using the Octet BLI platform, was developed to successfully assess the ability of antagonist candidates to inhibit the binding of VEGF to VEGF Receptor.
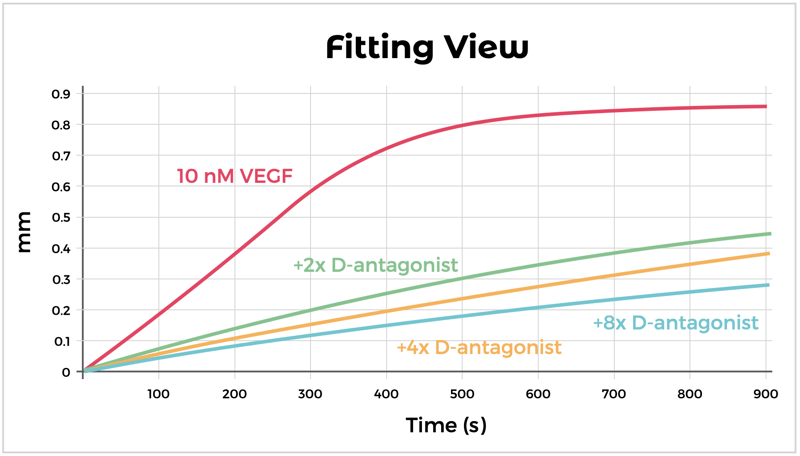
Using Bio-layer Interferometry to show inhibition of ligand binding (VEGF) to soluble receptor (VEGFR1) by a D-protein antagonist.
Going a step further, assays can be designed to measure the impact of antibody binding on a biological activity. The simplest approach is to use cells that express both a drug target and the bioactivity associated with the target. The bioactivity reported could be the activation or inhibition of a signaling pathway by way of measuring a downstream effect of signaling. For example, phosphorylation events after target binding can be monitored using phosphorylation-site specific antibodies, and promoter activation can be evaluated using reporter genes in engineered cell lines. Other assays for bioactivity induced by antibody binding or signaling include cell proliferation, cytotoxicity, antibody internalization, and migration. These assays may require a variety of custom reagents too extensive to be described here. Finally, complex bio-assays may need to be used to identify candidate therapeutic antibodies. One example, a natural killer activity assay, is shown below and in the associated poster.
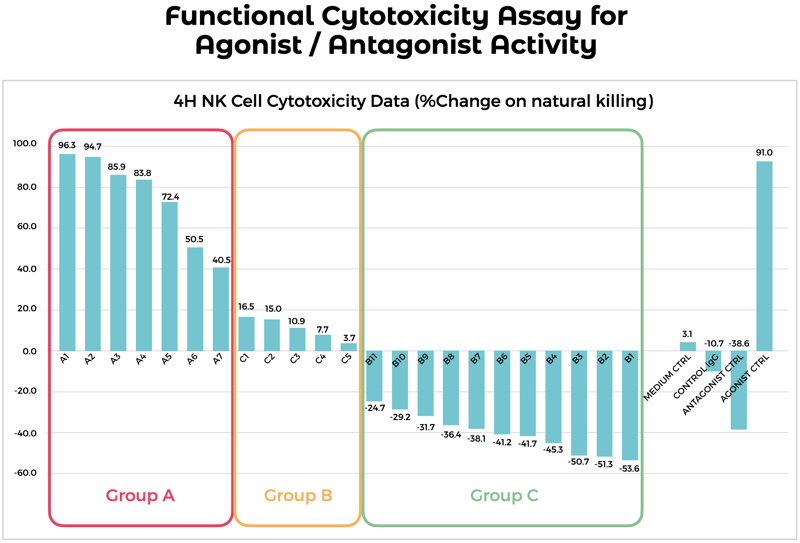
Using a Cytotoxicity Assay to Show Antibody Agonist or Antagonist Activity by Natural Killer Cells.
Using purified antibodies, a cytotoxicity assay was employed to determine the ability of candidate antibodies to act as either agonists or antagonists in the cell killing activity of Natural Killer (NK) cells. Antibodies raised against an immune checkpoint target expressed on NK cells were incubated with NK cells and target cancer cells. NK cell killing activity was monitored through fluorescent cell imaging. An increase in cancer cell death indicated antibodies which increased the activity of NK cells; a decrease in cancer cell death indicated antibodies that decreased NK cell activity. From this data set, both agonist and antagonist antibodies were identified for the modulation of NK cell activity for potential cancer therapeutics.
Having the right implements
In order to incorporate these types of biological activity assays into an antibody discovery campaign, particular tools and know-how are required. The table below shows a list of various functional assays and the necessary technology and equipment required to run them. Depending on the antigen, some of these assays can be easily incorporated into early screening efforts. The appropriate combination can quickly identify antibodies with binding and functional activity.
| ASSAY PLATFORM | Antigen Binding | Receptor Blocking | Cell Killing / Proliferation | Internalization Assay | Cell Signaling |
| ELISA | Yes | Yes | No | No | No |
| Flow Cytometry | Yes | Yes | Yes | Yes | Yes |
| BLI / SPR | Yes | Yes | No | No | No |
| Other (FRET / HTRF, etc) | Yes | Yes | Yes | No | Yes |
Strategic use of these tools can be a cost-effective and time-saving solution when expanding large numbers of candidate antibodies. Appropriate timing of assays is certainly a factor. Typically, antigen binding properties are first determined followed by functional assays. However, if antibodies with the desired function are expected to be rare, functional assays are best incorporated as early as possible. The type of antibody sample required (e.g., crude or purified) for an assay can impact the sequence of assays. Most binding assays can reliably be accomplished with serum or supernatant. By contrast, bioassays may require a purified antibody if other proteins, media components, or cell by-products are expected to impact the results.
Harvesting your crop of candidates
The ultimate goal of candidate identification is to find your “fit-for-purpose” antibodies. Efficiently and effectively evaluating the binding and functional activities of an antibody will enable you to reach this goal, and we’re eager to put our knowledge and skills to use toward your discovery project. Let’s start the conversation, and see what new crop of candidates we can harvest for you.

Josh is the Manager of Project Management of Antibody Solutions. Since joining the company in 2010, Josh has served multiple roles at the company and is currently providing scientific support and guidance to the Project Management team as well as helping clients achieve their antibody discovery and research needs.





